As the second aminoacyl tRNA binds to the A site, the initial tRNA molecule shifts to the adjacent P site. The A site is the first binding site where the initial incoming aminoacyl tRNA pairs with its codon. The three binding sites are the aminoacyl site (A site), peptidyl site (P site), and exit site (E site). There is more than one codon encrypting for each amino acid except methionine (Met) and tryptophan (Trp). A triplet of nucleotides identifies each amino acid, called a codon. Both ribosomal subunits join to form the binding sites for tRNA to translate the mRNA transcript. This creates protein isoforms when the downstream AUG is located in the same open reading frame as the first one, or a completely different protein if it is not in-frame.Įach ribosome is made up of a larger and smaller subunit, which is different for eukaryotes (the 60S and 40S) and prokaryotes (50S and 30S). The first AUG codon can be skipped (a process called “leaky scanning”) to use a downstream AUG. The PIC scans the mRNA in the 5’ region until it reaches the AUG start codon that is complementary with the anticodon of Met-tRNA. Ī favorable initiation sequence in mammals is known as “Kozak consensus”: 5’ (A/G)CCAUGG 3’. Several factors bind to this initiation sequence (IF4F initiation factors), forming the ribosome-eIF2-GTP-Met-tRNA preinitiation complex (PIC). A postulated mechanism (scanning mechanism) predicts that the ribosomes initiate protein synthesis recognizing and binding the 5′ tail of the mRNA corresponding to a 7-methylguanosine referred to as "cap." The first methionine codon located in the 5′ end of the mRNA is the initiator codon. 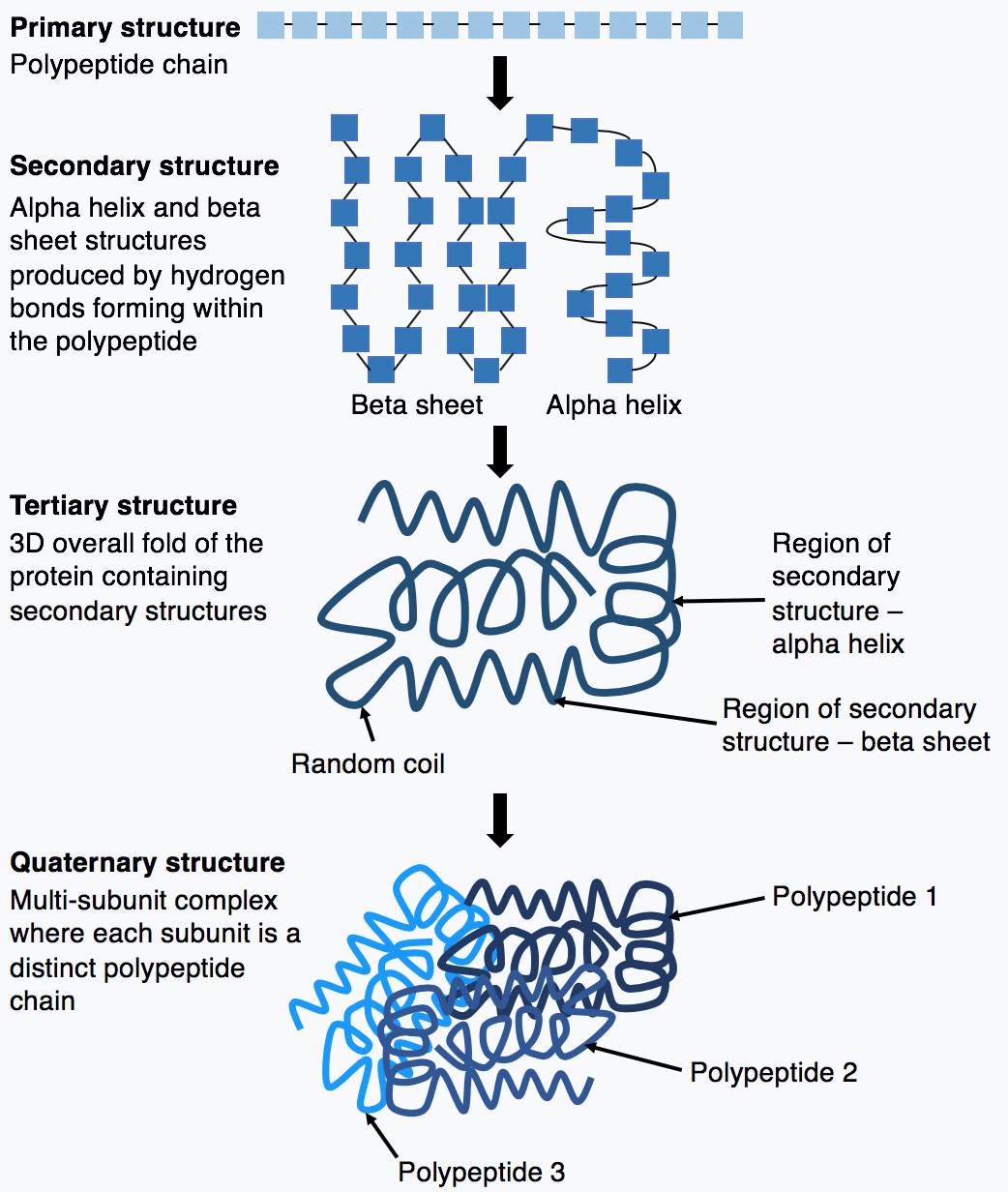
The initiation site in both prokaryotes and eukaryotes is not composed of a unique initiation sequence but is more probably a multisubstrate enzyme system based on substrate specificity. Īfter a mature mRNA transcript leaves the nucleus, a ribosome attaches to the transcript to begin translation. The mechanism underlying alternative splicing is not fully understood yet, but histone modification signatures correlate with splice site switching via chromatin-binding proteins. Alternative splicing is now known to deeply influence biological behavior (sex determination, cell differentiation, and programmed cell death). Alternative splicing is a mechanism used to create different isoforms of the same protein so that several protein isoforms can be created from a limited repertoire of genes. Splice sequences are found at the 5′ and 3′ tails of introns marked with the dinucleotide GU at its 5′ end and AG at the 3′ end. Exons are segments containing the encoding sequence for protein synthesis, while introns are intervening segments of nucleotides that will be excised from the primary RNA transcript (pre-mRNA) in a process called splicing. The transcribed RNA contains multiple segments called exons and introns. The consensus sequence TATTAA (called TATA box) is located 25–35 bases upstream of the initiation site, followed by a poly-G/C consensus sequence located 32–38 bases upstream. 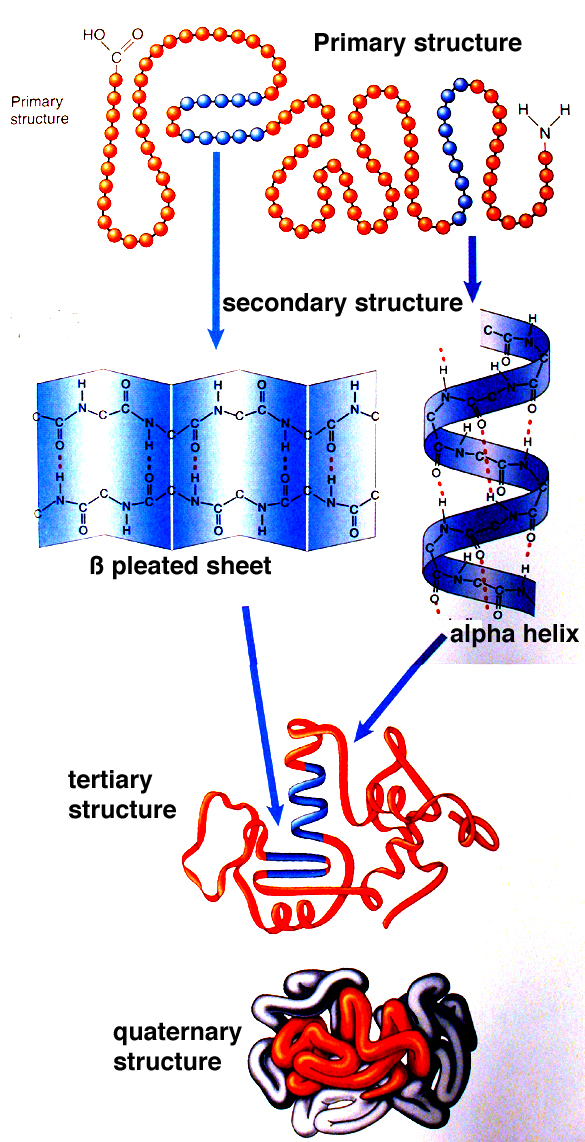
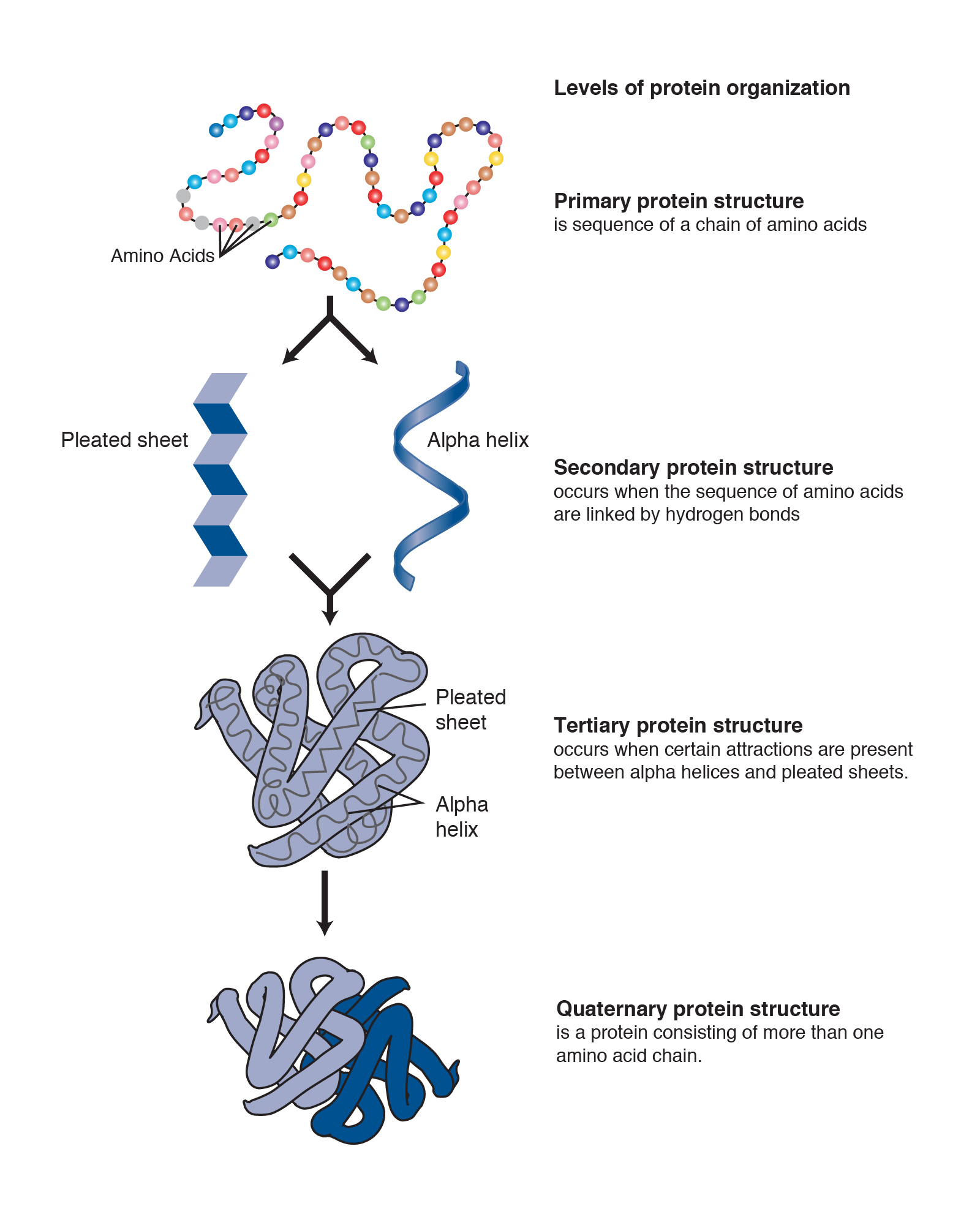
The initiation complex recognizes the promoter region in the DNA as a signal for the transcription starting point. The RNA polymerases are aided in the transcription function by a series of proteins called transcription factors, which have several functions mainly to facilitate the binding of RNA polymerase II to the promoter sequence, forming the transcription initiation complex. RNA polymerase III synthesizes 5S RNA, and all transfer RNA (tRNA) is located in the nucleoplasm. Only some proteins have a quaternary structure as well. Each protein at least contains a primary, secondary, and tertiary structure. The quaternary structure also influences the three-dimensional shape of the protein and is formed through the side-chain interactions between two or more polypeptides. The three-dimensional shape of a protein, its tertiary structure, is determined by the interactions of side chains from the polypeptide backbone. These hydrogen bonds create alpha-helix and beta-pleated sheets of the secondary structure. The secondary structure contains regions of amino acid chains that are stabilized by hydrogen bonds from the polypeptide backbone. 
Only some proteins have a quaternary structure as well. The primary structure is comprised of a linear chain of amino acids. Every protein at least contains a primary, secondary, and tertiary structure. A protein can be identified based on each level of its structure. They carry out a wide variety of organism functions, including DNA replication, transporting molecules, catalyzing metabolic reactions, and providing structural support to cells. Proteins are polypeptide structures consisting of one or more long chains of amino acid residues. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
